What is Hpath?
Hpath is an ontology of terms created based on INHAND code list and complemented to cover the needs of the ![]() eTOX project.
eTOX project.
An ontology aims at giving a hierarchical structure to preferred terms, in this case descriptions of histopathological findings.
Goal was to harmonize data manually curated from 6000+ regulatory toxicology study reports on 9 laboratory species, created within the last 40 years and donated by 13 companies. From these reports, 31 000+ terms were found describing histopathological alterations and additional 17 000+ terms were identified as describing the alteration location. These heterogeneous data did call for a normalization to a common ontology, which systematically classifies synonyms.
Synonyms were grouped and aligned with the INHAND terms from the available published manuscript. Additional terms where created to cover the missing manuscript and will be replaced by INHAND terms when they become available. The main advantage of an ontology compared to a code list is the presence of a tree structure linking terms according to the pathological process. It means that using parent nodes one can collect related findings.
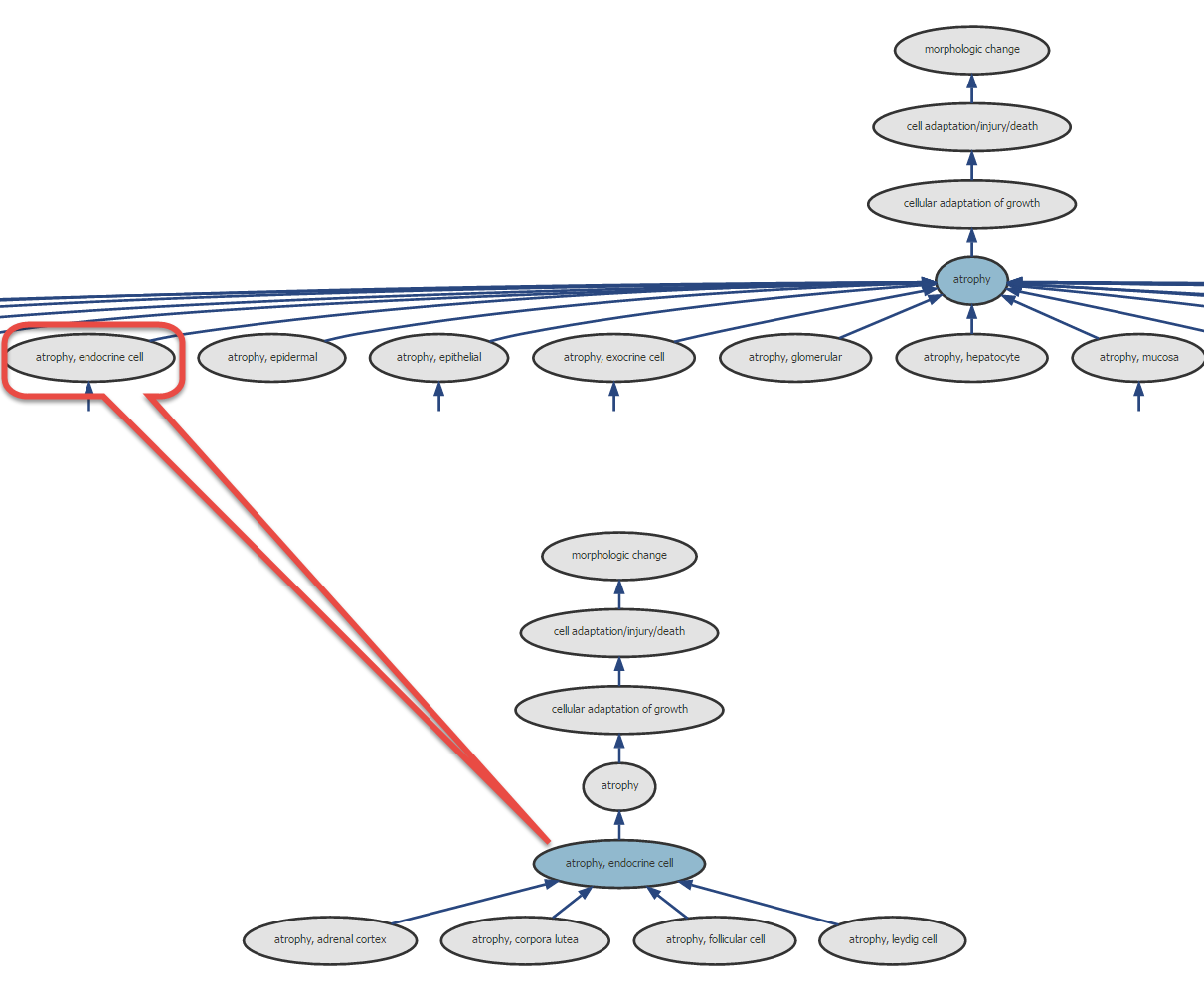
Figure 1: Example of term (e.g. the INHAND term “Atrophy, corpora lutea”), as a “child term”, a branching value from the parent node “atrophy, endocrine cell”. This latter term is further related to the above parent node “atrophy”, which also gathers many other different “child terms” from different organ/tissue.
Why is it useful?
Mining histo-pathology data is often difficult because of the variety of terms used, which often reflects difference in historical usage of the terms or focus on different aspects of a given pathological process, otherwise being often synonyms. Searches like historical finding occurrence or co-occurring finding patterns across studies are hard to conduct. In eTOX, the Hpath ontology was used to map term dictionaries and to make these data searchable both in eTOX web interfaces and using database query language.
The tree structure makes it easy for pathologists and toxicologists to explore the existing terms, identify relevant ones and their occurrence in vivo. It is also an important element for data mining, e.g. enabling the design of user search interfaces for toxicology data warehouses.
Electronic data submission using the Standard for Exchange of Non-Clinical Data (SEND) implemented by CDISC (Clinical Data Interchange Standards Consortium) will be enforced by the FDA for toxicology studies starting end of 2016. FDA has indicated a preference to utilize microscopic pathology terminology developed by INHAND. Therefore, INHAND terms will be mapped to the SEND terminology. Hpath could help the identification of the correct SEND term for all terms included.
How to use the ontology?
The use of the ontology is demonstrated by Figure 2, an example taken from the eTOX OntoBrowser interface. Let’s assume a user to be interested to learn more about the INHAND preferred term: “hyperplasia, squamous cell”. This term can be entered into the search field (right top). As results related terms are shown under the header search results, namely:
- "HYPERPLASIA SQUAMOUS CELL" EXACT [NIBR]
- "SQUAMOUS CELL HYPERPLASIA" EXACT [NIBR]
- "HYPERPLASIA, SQUAMOUS CELL" EXACT [NIBR]
- "HYPERPLASIA SQUAMOUS CELL" EXACT [NIBR]
- "Hyperplasia,Squam.C." RELATED [NIBR]
- "Squamous Hyperplasia" EXACT [NIBR]
- "Hyperplasia: squamous cell" EXACT [VX]
- "Hyperplasia, squamous cell" EXACT [NIBR]
- "Squamous cell hyperplasia" EXACT [NIBR]
- "Hyperplasia, squamous cell" EXACT [VX]
- "Hyperpl.,Squam.Cell" EXACT [NIBR]
- "Hyperplasia: squamous cell" EXACT [NIBR]
- "SQUAMOUS HYPERPLASIA" EXACT [NIBR]
- "Squamous hyperplasia" EXACT [VX]
- "Squamous cell hyperplasia" EXACT [VX]
- "Hyperpl. Squam.Cell" EXACT [NIBR]
- "Keratinized epithelial cells" RELATED [VX]
- "squamous cell hyperplasia" EXACT [VX]
- "HYPERPLASIA: SQUAMOUS CELL" EXACT [NIBR]
Further the user gets the information that all synonyms are allocated to one preferred term, which arises from INHAND. For a given term, the ontology describes both the cross references to INHAND + its definition, and the parent nodes of the hierarchical relation:
- xref: "INHAND:7022 "Hyperplasia, squamous cell"
- is_a: HPATH:0000083 ! hyperplasia
The tree structure, as depicted in the center of the Figure 1, informs the user about the hierarchical relation of terms, and the relation to other parent nodes in this ontology. In this case “hyperplasia, squamous cell” is a sub-term of “hyperplasia”. “Hyperplasia”, which in turn is a sub-term of “cellular adaptation of growth”, which is a sub-term of “cell adaptation/injury/death” and finally a sub-term of “morphological change”.
Parent nodes are also clickable, unfolding the full relation tree of the parent node in the ontology. This is shown in figure 2, a screenshot depicting sub-nodes of the term “cellular adaptation of growth”. “Hyperplasia” is 1 out of 10 sub-nodes. The arrow under the node indicates, where further sub-notes and thus more information is available.
This case exemplifies how the ontology is more than a collection of standardized vocabulary, by incorporating a lot of histopathological knowledge.
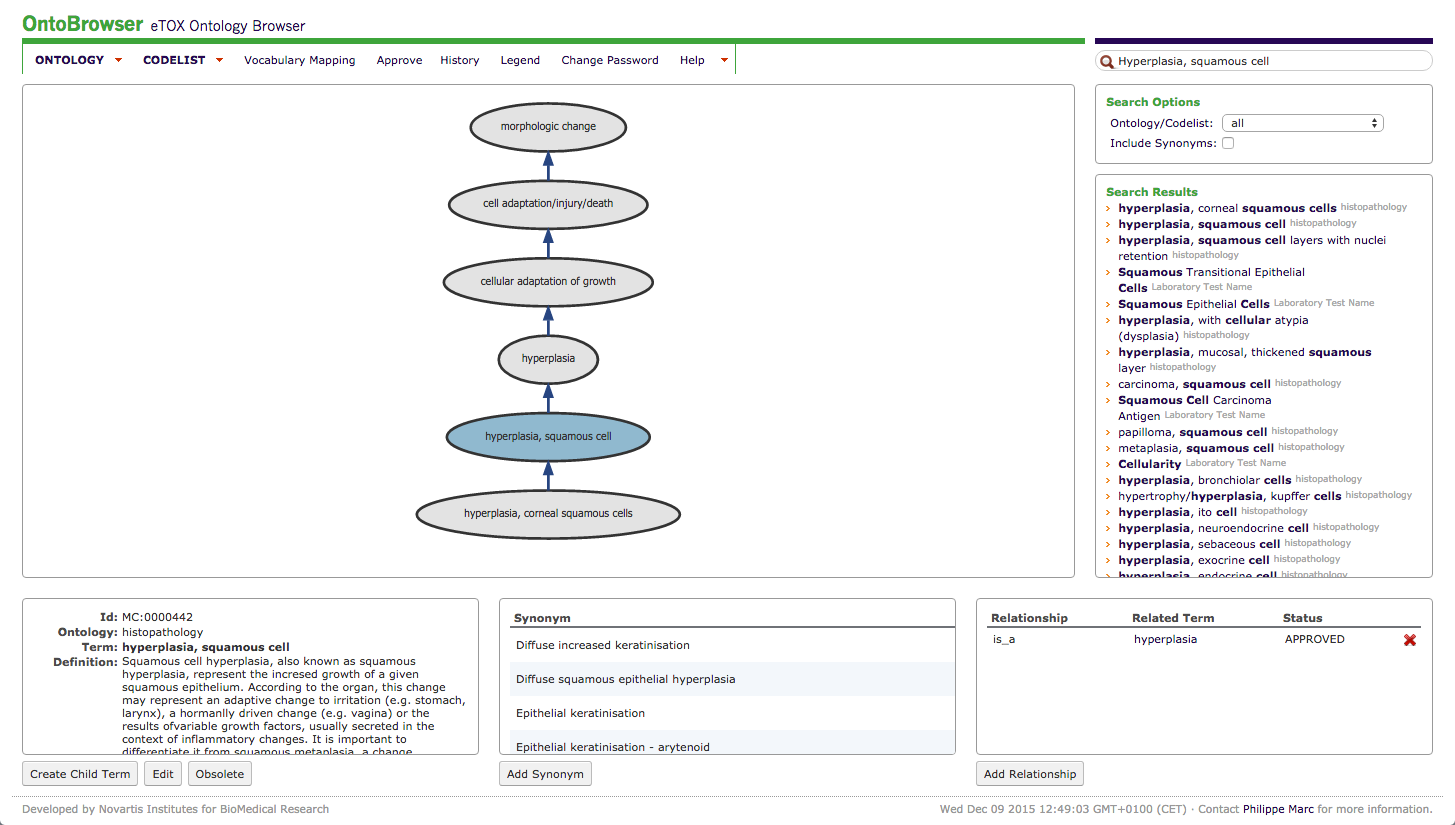
Figure 2: Screenshot from Ontobrowser: the results for the term “hyperplasia, squamous cell” are depicted.
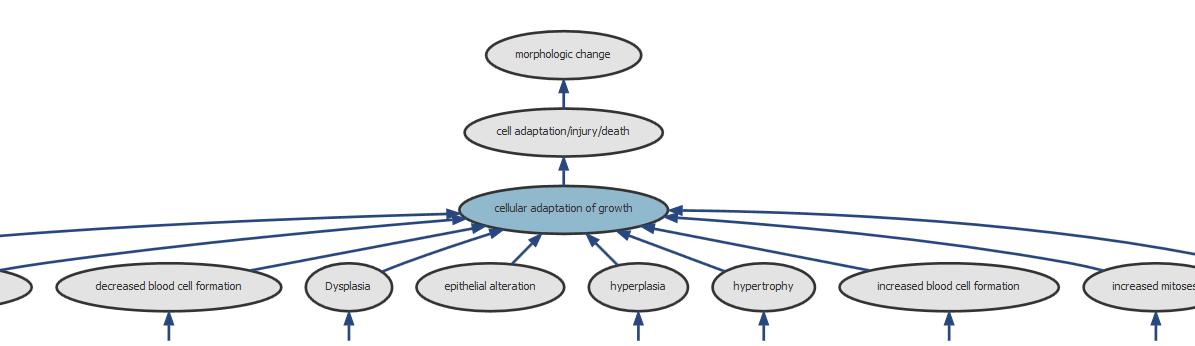
Figure 3: Screenshot of the information available for the parent node “cellular adaptation of growth”.
Who can use it?
The ontology will be released in 2016 under an open license (Apache License Version 2.0), giving unrestricted access to it for any usage and encouraging contribution.
Feedback welcome!
Feedback on the use, the applicability of the ontology to questions outside eTox will help to improve the ontology. Please feel free to contact: Alessandro Piaia (Alessandro.Piaia@Novartis.com)